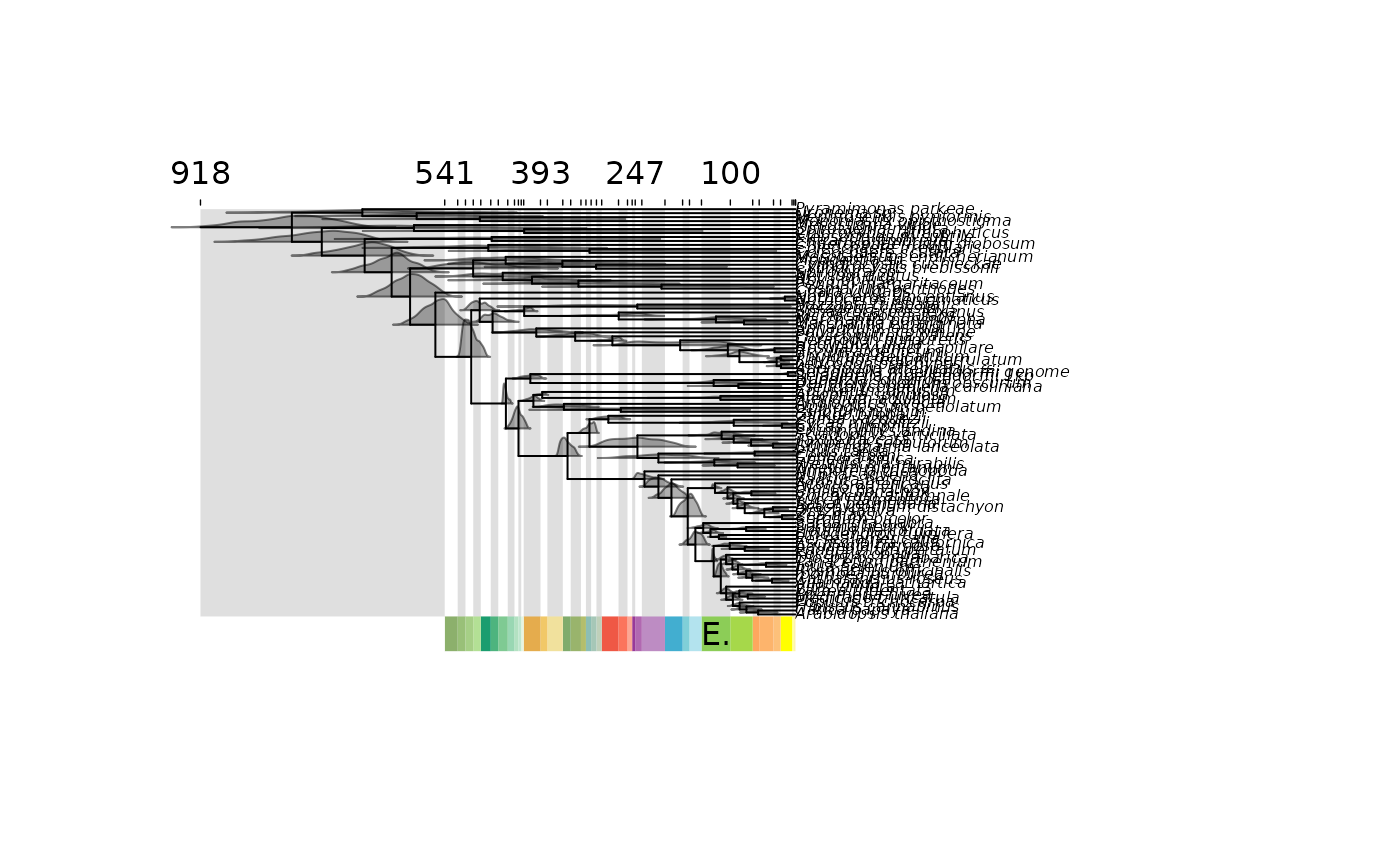Plot time-scaled phylogenies
mcmc.tree.plot.RdPlot time-scaled phylogenies with node uncertainty and timescale
Usage
MCMC.tree.plot(
phy = NULL,
analysis.type = "MCMCtree",
MCMC.chain = NULL,
node.ages = NULL,
directory.files = NULL,
plot.type = "phylogram",
build.tree = FALSE,
node.method = "bar",
all.nodes = NULL,
add.time.scale = TRUE,
add.abs.time = TRUE,
scale.res = "Epoch",
label.timescale.names = FALSE,
time.correction = 1,
col.age = "blue",
tip.lengths = FALSE,
density.col = "#00000050",
density.border.col = "#00000080",
cex.tips = 1,
show.tip.label = TRUE,
col.tree = "black",
tip.color = "black",
lwd.bar = 1,
grey.bars = TRUE,
cex.age = 1,
cex.labels = 1,
cex.names = 1,
relative.height = 0.08,
tip.bar.col = "#ff000050",
burn.in = 0.25,
distribution.height = 0.8,
abs.age.mgp = c(3, 0.35, 0),
abs.age.lwd.ticks = 0.7,
abs.age.lwd = 0,
tck.abs.age = -0.01,
abs.age.line = -0.4,
pos.age = NULL,
n.runs = 2,
ladderize.tree = TRUE,
...
)Arguments
- phy
A timescaled phylogeny, unless analysis.type="MCMCtree" and build.tree=TRUE
- analysis.type
The method used to generate the time-scale tree, one of MCMCtree, MrBayes, RevBayes, BEAST2, or User.
- MCMC.chain
The full posterior of age estimates for all nodes (default NULL)
- node.ages
List of user-supplied node ages applicable for analysis.type user. Either all nodes or a selection of nodes. Each list element must be named with its corresponding node label from the APE format.
- directory.files
The directory for files to summarise for MrBayes, RevBayes, and BEAST2 analyses
- plot.type
The plotting method for the phylogram corresponding to the APE definition. Phylogram is available for all analysis types, but cladogram is only avilable for MCMCtree analyses at present. Type distributions plots a phylogram with stats::density distributions on each of the nodes.
- build.tree
Logical. Only applicable to MCMCtree analyses, whether to timescale the phylogeny based on the full MCMC chain
- node.method
For plot.type phylogram the method to dispay age uncertainty on each node, either bar, node.length, or full.length. If 'none' is supplied plotting node uncertainty is suppressed.
- all.nodes
If NULL (default) node uncertainty is plotted on all nodes. If node numbers are supplied, only these nodes will be labelled with uncertainty.
- add.time.scale
Logical. Adds a timescale to the plotted phylogeny.
- add.abs.time
Logical. Adds an absolute timescale alongside the geological timescale. Only applicable if add.time.scale is TRUE
- scale.res
The geological age designation to add to the plot can be one or a combination of Eon, Period, Epoch, Age. The order of plot (from bottom to top) is the same as the supplied order. Subsequent arguments add.abs.time and grey.bars are based on the last supplied age designation.
- label.timescale.names
Logical. Add names (Eon, Period, Epoch, Age, Ma) to axis if timescale used
- time.correction
Number to place branch lengths and age estimates in absolute time, deafult to one.
- col.age
The colouring of the node.method bars to summarise node age uncertainty
- tip.lengths
Logical. If the tree contains non-present tip ages, the age uncertainty associated with them will be added to the plot
- density.col
Colour of the node distributions (if analysis.type=distributions)
- density.border.col
Colour of the node distributions (if analysis.type=distributions) borders
- cex.tips
Size of the phylogeny tip labels
- show.tip.label
Logical. Should the tree tip labels be displayed
- col.tree
Colour of the phylogeny edges
- tip.color
Colour of the phylogeny tip labels
- lwd.bar
Width of the bar to summarise age uncertainty, applicable only if plot.type is phylogram and node.method is bar
- grey.bars
Logical. Should grey bars be used to signify time bins, applicable only if add.time.scale is TRUE
- cex.age
Size of the labels for the absolute timescale
- cex.labels
Size of the labels for the geological timescale
- cex.names
Logical. Add label.timescale.names, if applicable.
- relative.height
the relative height of the timescale labels
- tip.bar.col
The colour of uncertainty around non-contemporary tips
- burn.in
The number of points in the chain to discard for MrBayes and RevBayes analyses
- distribution.height
The relative height of node distributions when plot.type is distributions measured as the relative height of the descendent node
- abs.age.mgp
mgp values for the absolute age axis, only applicable if add.abs.time is TRUE
- abs.age.lwd.ticks
lwd values for the absolute age axis ticks, only applicable if add.abs.time is TRUE
- abs.age.lwd
lwd values for the absolute age axis horizontal line, only applicable if add.abs.time is TRUE
- tck.abs.age
tck values for the absolute age axis tick height, only applicable if add.abs.time is TRUE
- abs.age.line
line correction for the absolute age axis tick, only applicable if add.abs.time is TRUE
- pos.age
position position of absolute age axis
- n.runs
For analysis.type mrbayes, the number of independent chains to summarise
- ladderize.tree
Logical. Plot a ladderized tree (TRUE) or not (FALSE)
- ...
further arguments to be used in
plot.phylo
Details
The primary inputs for the 'method' options are outputs from analysis conducting using MCMCtree, MrBayes, RevBayes, or User.
For analyses under the 'MCMCtree' option (the default) the function only requires a 'FigTree' output from MCMCtree analysis or the full MCMCtree posterior. For option "MrBayes" and "RevBayes", the method argument takes the user-supplied directory address in which all the output files are stored.
For the option 'User' the function requires a full posterior distribution and time-scaled phylogeny. This function can take any posterior distribution, but it may require some manipulation. The argument node.ages in MCMC.tree.plot takes a named list containing the posterior ages for each node, with the name of each element corresponding to the node in the phylogeny.
Examples
data(MCMCtree.output)
#> Warning: data set ‘MCMCtree.output’ not found
attach(MCMCtree.output)
## if it is necessary to read in file manually
## MCMCtree.posterior <- read.csv("mcmc.posterior.file.directory", sep="\t")
MCMCtree.file <- readMCMCtree(MCMCtree.phy, from.file=FALSE)$apePhy
MCMC.tree.plot(phy=MCMCtree.file, analysis.type="MCMCtree",
MCMC.chain=MCMCtree.posterior, plot.type="distributions", cex.tips=0.5,
time.correction=100)